Minimal Example¶
Most of the functionality of this package is bundled in two main classes: ESN and ESNWrapper. (ESN stands for for Echo State Network.) For all normal usage of the rescomp package, ESNWrapper is the class to use.
As always, we first want to import the needed packages.
[1]:
import rescomp
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
Then, we can create an instance of the ESNWrapper class to be used throughout this example.
[2]:
esn = rescomp.ESNWrapper()
As with most machine learning techniques, we need some training data, which we generate here artificially by simulating the chaotic Lorenz63 system for 5000 time steps.
[3]:
simulation_time_steps = 5000
[4]:
starting_point = np.array([-14.03020521, -20.88693127, 25.53545])
sim_data = rescomp.simulate_trajectory(
sys_flag='lorenz', dt=2e-2, time_steps=simulation_time_steps,
starting_point=starting_point)
As a convention, the data specified is always input data in an np.ndarray of the shape (T, d), where T is the number of time steps and d the system dimension I.g. RC works for any number of dimensions and data length.
Until now, the object esn is basically empty. One main component of reservoir computing is the reservoir, i.e. the internal network, wich we can create via create_network().
For the sake of consistency, we want to set the random seed used to create the random network, so that we always create the same network, and hence the same prediction, when running this example.
[5]:
np.random.seed(0)
Now we can create the network.
[6]:
esn.create_network()
The number of time steps used to synchronize the reservoir should be at least a couple hundred but no more than a couple thousand are needed, even for complex systems.
[7]:
train_sync_steps = 400
The number of time steps used to train the reservoir: This depends hugely on the system in question. See the FAQ for more details.
[8]:
train_steps = 4000
The number of time steps to be predicted:
[9]:
pred_steps = 500
Plug it all in:
[10]:
y_pred, y_test = esn.train_and_predict(
x_data=sim_data, train_sync_steps=train_sync_steps, train_steps=train_steps,
pred_steps=pred_steps)
The first output y_pred is the the predicted trajectory print(y_pred.shape)
If the input data is longer than the data used for synchronization and training, i.e. if
x_data.shape[0] > train_sync_steps + train_steps,
then the rest of the data can be used as test set to compare the prediction against. If the prediction where perfect y_pred and y_test would be the same Be careful though, if
x_data.shape[0] - (train_sync_steps + train_steps) < pred_steps
then the test data set is shorter than the predicted one:
y_test.shape[0] < y_pred.shape[0].
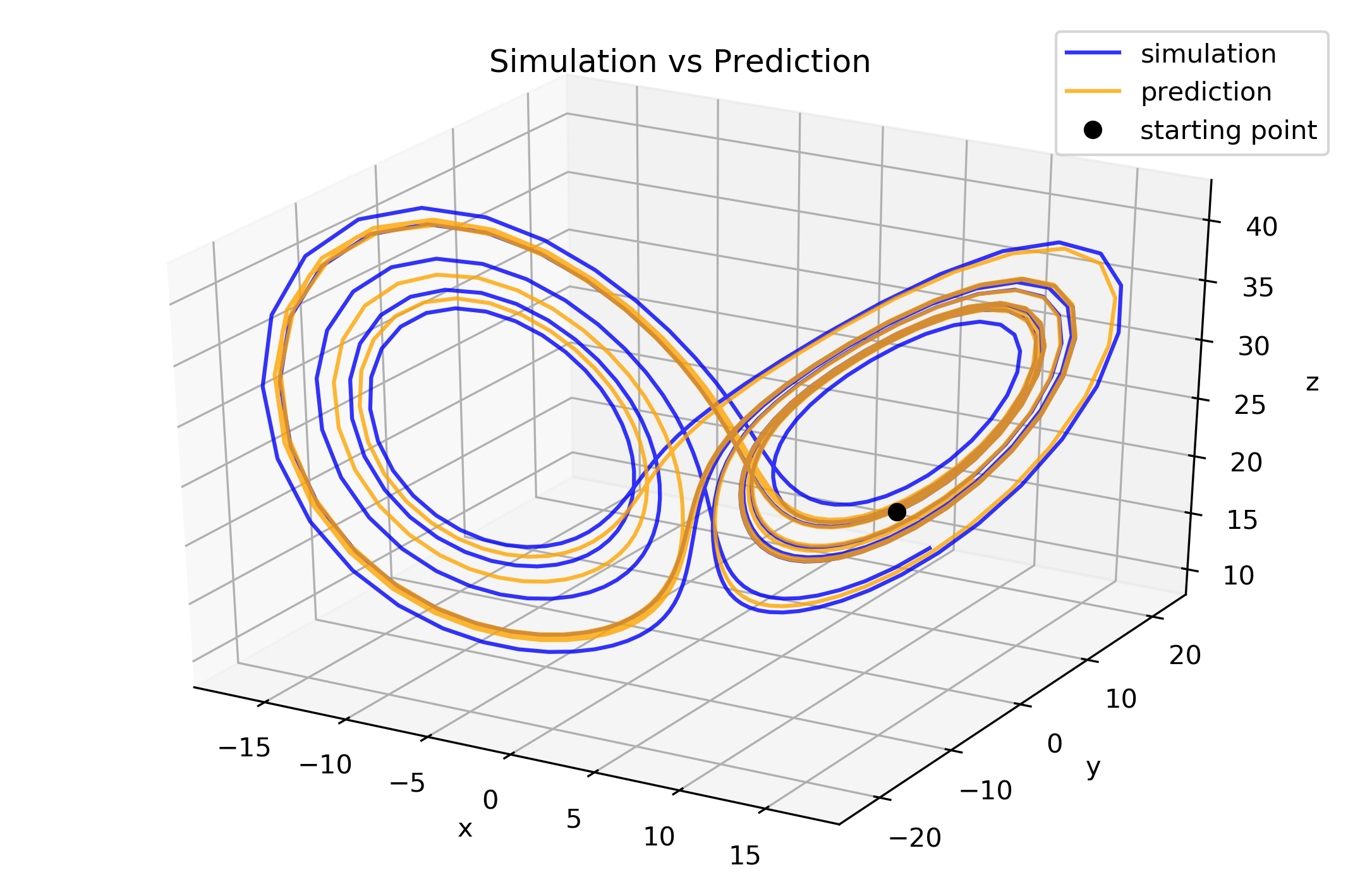
Plot prediction and simulation to compare them
[11]:
fig1 = plt.figure(figsize=(9, 6), dpi=300)
ax1 = fig1.add_subplot(111, projection='3d')
ax1.plot(y_test[:, 0], y_test[:, 1], y_test[:, 2],
alpha=0.8, color='blue', label='simulation')
ax1.plot(y_pred[:, 0], y_pred[:, 1], y_pred[:, 2],
alpha=0.8, color='orange', label='prediction')
start = y_pred[0]
ax1.plot([start[0]], [start[1]], [start[2]],
color='black', linestyle='', marker='o', label='starting point')
ax1.set_title("Simulation vs Prediction")
ax1.set_xlabel('x')
ax1.set_ylabel('y')
ax1.set_zlabel('z')
plt.legend()
plt.show()
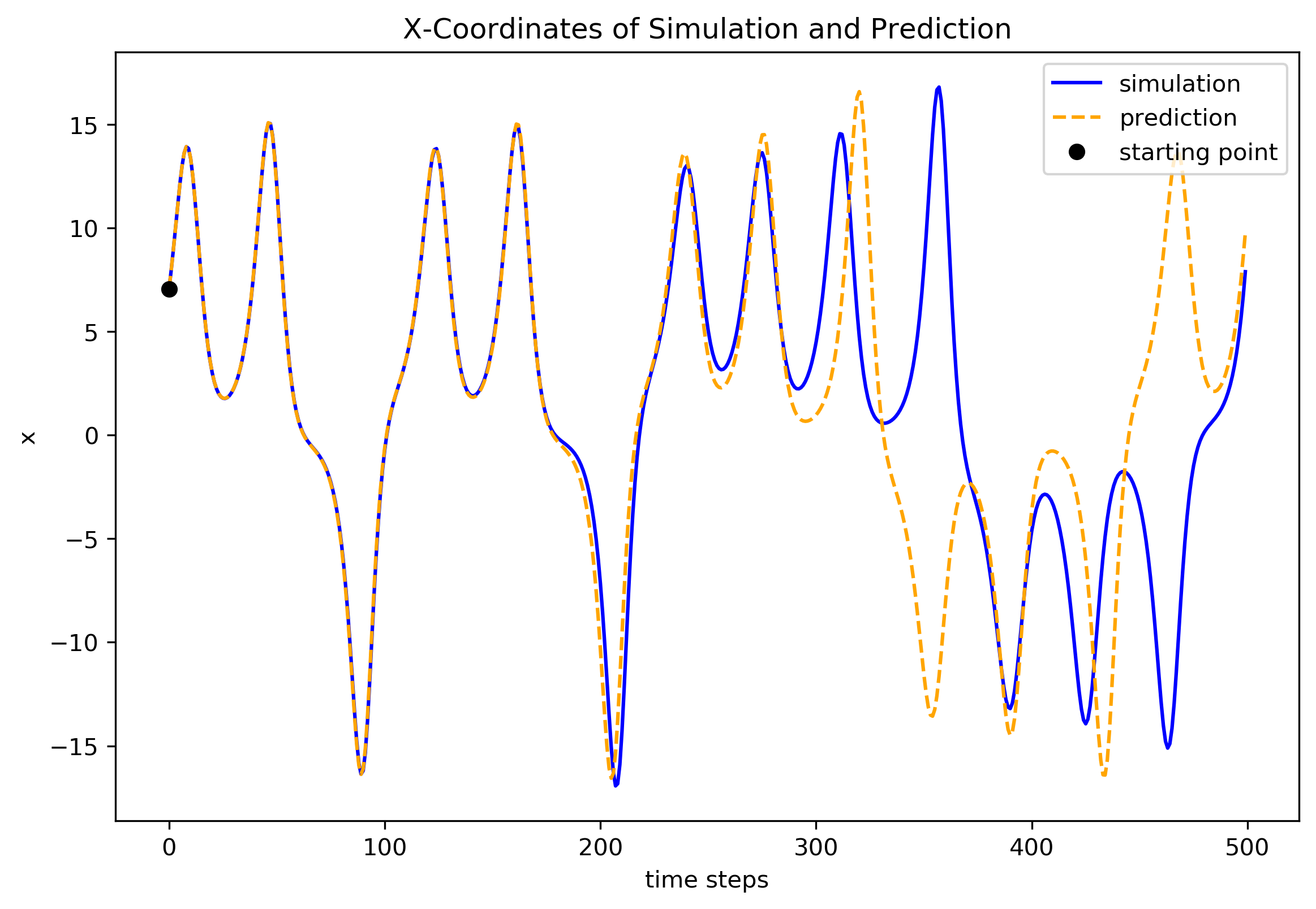
Now, plot their x coordinates as a more detailed comparison.
[12]:
fig2 = plt.figure(figsize=(9, 6), dpi=300)
ax2 = fig2.add_subplot(1, 1, 1)
ax2.plot(y_test[:, 0], color='blue', label='simulation')
ax2.plot(y_pred[:, 0], color='orange', linestyle='--', label='prediction')
start = y_pred[0]
ax2.plot(start[0], color='black', linestyle='', marker='o',
label='starting point')
ax2.set_title("X-Coordinates of Simulation and Prediction")
ax2.set_xlabel('time steps')
ax2.set_ylabel('x')
plt.legend()
plt.show()